
(*** 8/22/23: We are currently recruiting lab techs and/or managers. For more info, contact: j.taylor60@csuohio.edu ***)
In the Taylor Lab, our goal is to better understand the relationship between epigenetic modifications and the aging process. Epigenetic alterations that occur during aging are a primary driver of age-related dysfunction and disease. Remarkably, interventions that target the epigenome can reverse age-related impairments and extend lifespan in laboratory animals.
Our lab currently uses Drosophila melanogaster (fruit flies) to study (i) how the epigenome changes with age and disease, and (ii) how experimental manipulation of the epigenome affects health, longevity, and the progression of various human disease models (such as Alzheimer’s disease). By studying these questions, our goal is to develop strategies to help reduce disease and disability in humans as they age. We employ a variety of techniques, including next-generation sequencing and bioinformatic analysis, genetic engineering, and population longevity experiments.
Most recently, we discovered that increasing levels of the epigenetic-modifier gene Sirt6 extends lifespan and preserves physical activity with age in flies. This pro-longevity effect of increased Sirt6 levels is mediated through enhanced deacetylation of Myc target genes, leading to their transcriptional repression, which in turn leads to decreased protein synthesis – a phenotype associated with slowed aging. Currently our lab is focused on identifying additional molecular and tissue-specific mechanisms by which Sirt6 regulates aging, and exploring the potential role of Sirt6 in Alzheimer’s Disease. We are also performing screens to identify new epigenetic modifiers of aging.
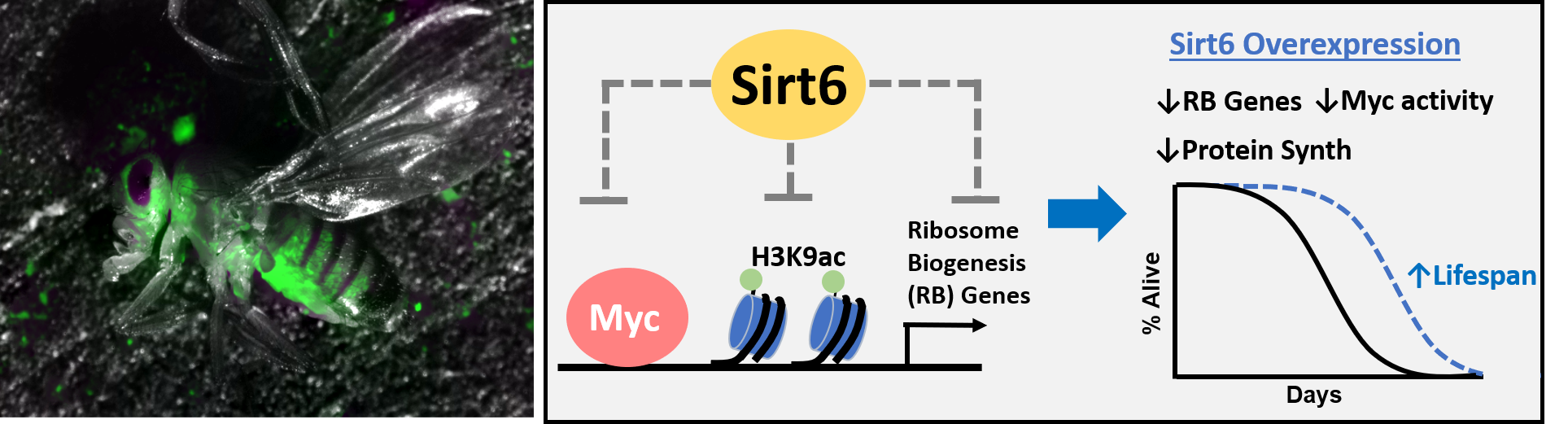
(Left)Transgenic fruit fly engineered to express GFP when epigenetic perturbations occur. (Right) Summary mechanism for lifespan extension by Sirt6 overexpression. Sirt6 overexpression epigenetically represses Myc target genes involved in ribosome biogenesis, by removing the activating histone mark H3K9ac in the TSS/proximal promoter region, leading to reduced protein synthesis and lifespan extension.
The Taylor lab was established at Cleveland State University in Fall, 2022. Dr. Taylor received his Ph.D. in Neuorscience from Wake Forest University School of Medicine in 2013, and performed his postdoctoral training at Wake Forest Health Sciences (2013-2016) and Brown Univeristy (2016-2022).
Selected Publications::
Taylor JR, Wood JW, Chang CC, Mizerak EM, Finn M, Liu JL, Hinthorne SL, Gordon S, Hutfilz CR, Klein MA, Denu JM, Gorbunova V, Boeke JD, Sedivy JM, Helfand SL. Sirt6 regulates lifespan in Drosophila melanogaster. Proceedings of the National Academy of Sciences, 2022
Gorbunova V, Seluanov A, Mita P, Boeke JD, Linker S, Gage FH, Kreiling JA, Petrashen AP, Taylor JR, Helfand SL, Sedivy JM. The involvement of transposable elements in aging and age-associated diseases. Nature, 2021
Taylor JR, Reynolds L, Hou L, Lohman K, Cui W, Kritchevsky S, McCall C, Liu Y. Transcriptomic profiles of aging in naïve and memory CD4+ cells from mice. Immunity & Ageing, 2017
Reynolds LM*, Taylor JR*, Ding J, Lohman K, Johnson C, Siscovick D, Burke G, Post W, Shea S, Jacobs DR Jr., Stunnenberg H, Kritchevsky SB, Hoeschele I, McCall CE, Herrington D, Tracy RP, Liu Y. Age-related variations in the methylome associated with gene expression in human monocytes and T cells. Nature Communications 2014 *co-first authors
Taylor JR, Zhang T, Messi ML, Wang ZM, Hereñú C, Kuan PF, Delbono O. (2014) The Cavβ1a Subunit Regulates Gene Expression and Suppresses Myogenin in Muscle Progenitor Cells. J Cell Biol., 205(6): 829-846.
(Last updated: 8/22/23)


